Scientists are striving to develop an immunotherapy targeting proteins resulting from mutations in cancer-driving genes. One team is now a step closer: they have identified a tumor-specific target derived from a commonly mutated driver gene, as well as a T cell receptor (TCR) that recognizes and binds it. Using the U.S. Department of Energy’s Advanced Photon Source (APS) for macromolecular x-ray crystallography studies, the researchers uncovered the mechanism behind the mutated protein’s immunogenicity, or ability to generate an immune response. The x-ray crystallography was part of a multipronged strategy that included a high-throughput T cell discovery platform, in vivo detection of circulating T cells in cancer patients, and adoptive cell transfer in a xenograft mouse model. The research, published in the journal Nature Medicine, provides insights into developing a targeted therapy that fights a variety of cancers, in multiple patients, at the cancer’s source.
In the past 20 years, neoantigens have become the holy grail of cancer immunotherapy. A neoantigen is a protein produced by mutations in the cell’s DNA. Private neoantigens result from the many “passenger” mutations that occur within a tumor environment but don’t contribute to the fitness of the tumor. They are patient-specific and therefore require customized therapeutic products. Public neoantigens arise from mutations in genes that are critical to cancer formation and maintenance. A key feature of public neoantigens is that they are compatible with common tissue types, also known as HLA. Because they are frequently conserved across patient populations regardless of what type of cancer they have, such a therapeutic approach has the potential for wider clinical application.
However, immunogenic public neoantigens are incredibly difficult to find, as they depend on the perfect alignment of three highly variable factors: 1) the cancer cell efficiently processing the mutated protein for presentation; 2) the ability for a degraded protein, known as a peptide, to stably bind to the “correct” version of the HLA molecule and expressed on the cell surface; and 3) the peptide-HLA complex stimulating a T cell response.
The team chose the PIK3CA gene as their proof-of-concept tumor target. It is one of the most frequently mutated driver oncogenes across many cancers, including breast, endometrial, ovarian, and bladder cancers. To explore whether PIK3CA could generate an immunogenic public neoantigen, the team employed a variety of techniques, including high-throughput single-cell transcriptomics, T cell receptor (TCR) sequencing, and liquid chromatography–tandem mass spectrometry. These technologies allowed them to identify which variation (allele) of HLA molecule was involved, as well as the amino acid sequence of the peptide and a candidate TCR.
Importantly, the scientists wanted to shed light on the mechanism of immunogenicity of mutated PIK3CA. For this, they turned to x-ray crystallography and the APS, an Office of Science user facility at Argonne National Laboratory.
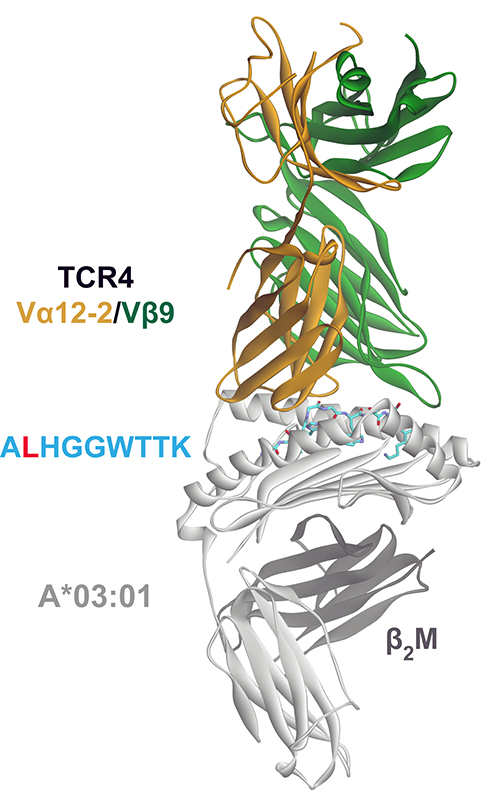
The team’s crystallographers prepared two different sets of materials: the normal (wildtype) or the mutated neopeptides bound to the HLA allele, and the peptide-HLA complex bound to a lead clinical T cell receptor candidate. After sending the materials to the staffs of the Southeast Regional Collaborative Access Team (SER-CAT) and the Northeastern Collaborative Access Team (NE-CAT) at the APS, journal article co-author Jiaqi Ma used remote control to illuminate the crystals from afar on SER-CAT x-ray beamline 22-ID and NE-CAT beamline 24-ID-E at 2.0Å and 3.1Å resolution, respectively. The diffraction data (Fig. 1) were downloaded back to the researchers’ home institutions from the remote-control system for analysis.
Based on the crystal structures, the team discovered that the replacement of a single amino acid in the mutated peptide enabled it to bind to the HLA molecule, relative to its wildtype counterpart, which was found to be highly unstable even at physiological temperature. The substitution also let the peptide-HLA molecule reside stably on the cell surface for an extended period of time, a property that has been correlated with the immunogenicity of a peptide-HLA complex. These studies were conducted using differential scanning fluorimetry and fluorescence anisotropy assays.
Analyzing the crystal structures of the peptide-HLA complex bound to the TCR, the scientists found that the receptor possessed an unusually long “peptide-sensing CDR3” loop. The extended loop allowed the receptor to have extensive interactions along the length of the neopeptide, giving it high affinity and very fine specificity.
The team wanted to establish the ability of mutated PIK3CA-specific T cell receptors to therapeutically engage cancer cells. They collected human blood samples and found evidence of circulating PIK3CA public neoantigen-specific T cells in cancer patients, confirming that the neoantigen stimulated an immune response.
In order to test the efficacy of a lead clinical candidate to mediate tumor regression in vivo, the researchers established a xenograft mouse model. They implanted mice with a tumor expressing the public neoantigen and transferred T cells engineered with the TCR. The scientists were thrilled to see very clear evidence of tumor control by the engineered T cells.
This is the basis of an immunotherapy treatment called adoptive cell transfer, in which T cells are taken from a patient, engineered in a lab to target the cancer cells, and reintroduced into the patient.
Based on the success of their findings in this research, the scientists are focusing on the clinical development of this immunotherapeutic strategy to be tested in clinical trials. ― Judy Meyers
See: Smita S. Chandran1,2*, Jiaqi Ma3, Martin G. Klatt1, Friederike Dündar4, Chaitanya Bandlamudi1, Pedram Razavi1,4, Hannah Y. Wen1, Britta Weigelt1, Paul Zumbo4, Si Ning Fu1, Lauren B. Banks1, Fei Yi1, Enric Vercher1, Inaki Etxeberria1, Watchain D. Bestman1, Arnaud Da Cruz Paula1, Ilinca S. Aricescu1, Alexander Drilon1,4, Doron Betel4, David A. Scheinberg1, Brian M. Baker3, and Christopher A. Klebanoff1,2,4**, “Immunogenicity and therapeutic targeting of a public neoantigen derived from mutated PIK3CA,” Nat. Med. 28, 946 (May 2022). DOI: 10.1038/s41591-022-01786-3
Author affiliations: 1Memorial Sloan Kettering Cancer Center, 2Parker Institute for Cancer Immunotherapy, 3University of Notre Dame, 4Weill Cornell Medicine
Correspondence: * [email protected], ** [email protected]
This study was supported, in part, by National Institutes of Heath (NIH) grants R37CA259177 (C.A.K.), P30CA008748 (C.A.K. and D.A.S.) and R01AI129543 (B.M.B.); the Parker Institute for Cancer Immunotherapy (C.A.K.); the Damon Runyon Cancer Research Foundation CI-96-18 (C.A.K.); Cancer Research Institute CRI3176 (C.A.K); the Functional Genomics Initiative Rapid Response Pilot Grant (C.A.K.); the Manhasset Women’s Coalition Against Breast Cancer (C.A.K.); the Emerson Collective Cancer Fund (C.A.K.); the Breast Cancer Alliance Exceptional Project Grant (C.A.K.); the Sussman Family Fund (C.A.K.); and a sponsored research agreement with Intima Bioscience (C.A.K.). SER-CAT is supported by its member institutions, and equipment grants (S10_RR25528, S10_RR028976 and S10_OD027000) from the National Institutes of Health. NE-CAT is funded by the National Institute of General Medical Sciences from the National Institutes of Health (P30 GM124165). The Eiger 16M detector on the 24-ID-E beam line is funded by a NIH-ORIP HEI grant (S10OD021527). M.G.K. was supported by an individual research grant from the German Research Foundation (DFG, KL3118/1-1). L.B.B. was supported, in part, by the Conquer Cancer Foundation. E.V. was supported by Programa de Ayudas de Movilidad para la Obtención de Mención ‘Doctor Internacional’ of the University of Navarra, FUNDACIONCAJANAVARRA-Obra Social ‘La Caixa’ and the Company of Biologists Travelling Fellowships Grant. F.Y. was supported, in part, by a fellowship from the Hearst Foundation. I.X. was supported, in part, by a fellowship from the Fundación Alfonso Martín Escudero and the Parker Institute for Cancer Immunotherapy. This research used resources of the Advanced Photon Source, a U.S. DOE Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
The U.S. Department of Energy's APS at Argonne National Laboratory is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
