The original Argonne press release by Emily Velasco and Andre Salles can be read here.
The nucleus of every cell in our bodies contains many complicated parts. One of the most intricate is called the nuclear pore complex, or NPC. It’s made up of more than 1,000 individual proteins and serves as a discriminating gatekeeper for the cell’s nucleus, the membrane-bound region inside a cell that holds its genetic material. Anything going in or out of the nucleus has to pass through the NPC on its way.
Understanding how the NPC works could have major implications for healthcare research. Not only is the NPC central to the operations of the cell, it is also involved in a variety of diseases. Mutations in the NPC are responsible for some incurable cancers, neurodegenerative and autoimmune diseases, such as amyotrophic lateral sclerosis (better known as Lou Gehrig’s disease, or ALS), and heart conditions including atrial fibrillation and early sudden cardiac death. Additionally, many viruses, including the one responsible for COVID-19, target and shut down the NPC during the course of their lifecycles.
André Hoelz, professor of chemistry and biochemistry at Caltech, Howard Hughes Medical Institute Faculty Scholar, and Heritage Medical Research Institute Investigator, has spent more than 20 years studying and deciphering the structure and function of the NPC, making extensive use of the Advanced Photon Source (APS), a U.S. Department of Energy (DOE) Office of Science user facility located at DOE’s Argonne National Laboratory.
Now, in a pair of papers published in the journal Science, “Architecture of the cytoplasmic face of the nuclear pore” [1] and “Architecture of the linker-scaffold in the nuclear pore,” [2][ Hoelz and his research team have unveiled the results of their work: a full structural map of the outer face of the NPC and a deeper understanding of the mechanism by which special proteins act like a molecular glue to hold the NPC together. This monumental result is the culmination of nearly 20 years of using ultrabright x-ray beams to illuminate the structure of the NPC, protein by protein.
For much of that time, Hoelz and his colleagues have used the National Institute of General Medical Sciences and National Cancer Institute (GM/CA-XSD) structural biology beamlines at the APS. Hoelz and his team determined their first structure at GM/CA-XSD in 2008 and have had a regular presence at the beamlines ever since.
“The staff at GM/CA-XSD realized in the beginning that what we were doing was not just one structure, but something major that required a long commitment,” Hoelz said. “They embraced from the get-go advances that helped with challenging samples. We tried every trick in the book to get these structures. Without access to high-end facilities such as the APS, we could not have done this.”
The researchers also made use of a Northeast Collaborative Access Team beamline at the APS early in their study. They also used the Stanford Synchrotron Radiation Lightsource (SSRL) at DOE’s SLAC National Accelerator Laboratory and the National Synchrotron Light Source II at DOE’s Brookhaven National Laboratory.
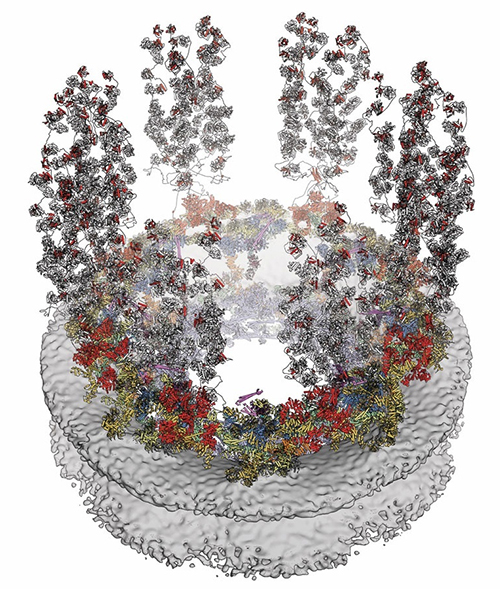
While the NPC can generally be thought of as akin to a door, it is unlike any you are going to encounter in your everyday life. Aside from looking like a donut with spider legs, the NPC doesn’t so much open and close as it does expand and contract.
“It’s a little like an airplane hangar where you can repair 747s, and the door opens to let the 747 come in, but there’s a person standing there who can keep a single marble from getting out while the doors are open,” Hoelz said.
To map the structure of this complex piece of machinery, Hoelz and his colleagues had to solve the equivalent of a very tiny 3-D jigsaw puzzle, using electron microscopy and x-ray crystallography on each puzzle piece. They began by growing the proteins that make up the NPC, and then testing them in pairs to see which ones fit together. If two puzzle pieces were found that fit together, the researchers would test the now-combined proteins against a third until they found the next piece, and so on.
Working their way through the proteins in this way eventually produced the final result of their first paper: the 16-protein wedge that gets repeated eight times around the face of the NPC like slices of a pizza.
This result complemented research conducted by Martin Beck of the Max Planck Institute of Biophysics in Frankfurt, Germany, whose team generated a map that provided the contours of a puzzle that the researchers placed the pieces into. To accelerate the completion of the puzzle of the human NPC structure, Hoelz and Beck exchanged data more than two years ago, and then independently built structures of the entire NPC.
“The substantially improved Beck map showed much more clearly where each piece of the NPC — for which we determined the atomic structures — had to be placed,” Hoelz says. “It was much akin to a wooden frame that defines the edge of a puzzle.”
The team used a similar process to determine the entire structure of what is known as the NPC’s linker-scaffold. This is a collection of proteins that help hold the NPC together while also providing it with the flexibility it needs to open and close, and to adjust to fit the molecules that pass through.
Hoelz likens the NPC to something built out of Lego bricks, but ones that do not always lock together. Instead they are lashed together by rubber bands that keep them mostly in place while still allowing them to move around a bit.
“I call these unstructured glue pieces the ‘dark matter of the pore,’ ” Hoelz says. “This elegant arrangement of spaghetti noodles holds everything together.”
At the APS, Hoelz and his team used the x-ray crystallography technique to determine dozens of these structures. The technique involves growing crystals from proteins, and then shining bright x-ray light onto those crystals. When the light diffracts or is scattered in different directions, a detector records the position and intensity of the light, and from that data scientists can glean important information about the atoms that make up the proteins.
Hoelz began working at GM/CA-XSD in 2008, connecting with Argonne’s Bob Fischetti, GM/CA-XSD group leader and life sciences advisor to the APS director. One feature of the beamline that made Hoelz’s work possible was the ability to change the size of the x-ray beam, down to incredibly small spots. This allowed Hoelz to work with smaller crystals, or crystals that did not diffract as well as others, he said.
“If you have a big crystal that diffracts well, it’s best to illuminate the whole crystal,” Fischetti said. “But if you have a smaller one, or a larger one that doesn’t diffract well, you can use a smaller beam to find the parts of the crystal that will give you the data you need.”
Fischetti and his colleagues won an R&D 100 award in 2010 for designing a system that allows scientists to choose the size of the x-ray beam. He called Hoelz’s work “phenomenal,” noting that because of his persistence, we now have a clear picture of one of the largest structures in the human cell.
“Our work at GM/CA would have been impossible without the people who do this day to day,” Hoelz said. “They try to help you in any way possible, and that’s an important thing. The people who work there are highly-trained scientists, and they’re the unsung heroes of all the research that is done there.”
Taken together, the findings of the two papers represent a leap forward in scientists’ understanding of how the NPC is built, and how it works. Caltech’s Christopher Bley, senior postdoctoral scholar research associate in chemistry and an author on both papers, says the team’s discoveries open the door for much more research.
“We built a framework on which a lot of experiments can now be done,” Bley said. “We have this composite structure now, and it enables and informs future experiments on NPC function, or even diseases. There are a lot of mutations in the NPC that are associated with terrible diseases. Knowing where they are in the structure and how they come together may help design the next set of experiments to try and answer the questions of what these mutations are doing.”
Hoelz also sees it as a testament to the value of empirical research in structural biology. While predictions can be helpful, he said, there’s no substitute for doing the work and determining the structures.
“There’s no ambiguity when structures are experimentally determined,” he said. “It’s hard data. Certainty is certainty.”
[1] See: Christopher J. Bley1, Si Nie1, George W. Mobbs1, Stefan Petrovic1, Anna T. Gres1,Xiaoyu Liu1, Somnath Mukherjee2, Sho Harvey1, Ferdinand M. Huber1, Daniel H. Lin1, Bonnie Brown1, Aaron W. Tang1, Emily J. Rundlet1, Ana R. Correia1, Shane Chen3,Saroj G. Regmi3, Taylor A. Stevens1, Claudia A. Jette1, Mary Dasso3, Alina Patke1, Alexander F. Palazzo4, Anthony A. Kossiakoff2, André Hoelz1*, “Architecture of the cytoplasmic face of the nuclear pore,” Science 376, eabm9129 (10 June 2022). DOI: 10.1126/science.abm9129
Author affiliations: 1California Institute of Technology, 2The University of Chicago, 3National Institutes of Health, 4University of Toronto
Correspondence: * [email protected]
The Molecular Observatory at the California Institute of Technology (Caltech) is supported by Donald and Judith Voet, the Gordon and Betty Moore Foundation, and the Beckman Institute. The operations at the SSRL are supported by the US Department of Energy and the National Institutes of Health (NIH). GM/CA-XSD has been funded in whole or in part with federal funds from the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006). S.P. and F.M.H. were supported by a PhD fellowship of the Boehringer Ingelheim Fonds. D.H.L. was supported by an NIH Research Service Award (5 T32 GM07616) and Amgen Graduate Fellowship through the Caltech-Amgen Research Collaboration. S.C., S.G.R., and M.D. were supported by National Institute of Child Health and Human Development Division of Intramural Research funding ZIAHD001902 and ZIAHD008954. A.A.K. was supported by NIH grants R01-GM117372 and P50-GM082545. A.H. was supported by a Camille-Dreyfus Teacher Scholar Award (TC-15-082) and NIH grants R01-GM117360 and R01-GM111461 and is an Investigator of the Heritage Medical Research Institute (HMRI-15-09-01) and a Faculty Scholar of the Howard Hughes Medical Institute (55108534). This research used resources of the Advanced Photon Source, a U.S. DOE Office of Science user facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357
[2] See: Stefan Petrovic, Dipanjan Samanta, Thibaud Perriches, Christopher J. Bley, Karsten Thierbach, Bonnie Brown, Si Nie, George W. Mobbs, Taylor A. Stevens, Xiaoyu Liu, Giovani Pinton Tomaleri, Lucas Schaus, and André Hoelz*, “Architecture of the linker-scaffold in the nuclear pore,” Science 376, eabm9798 (10 June 2022). DOI: 10.1126/science.abm9798
Author affiliation: California Institute of Technology
Correspondence: * [email protected]
The Molecular Observatory at Caltech is supported by D. and J. Voet, the Gordon and Betty Moore Foundation, and the Beckman Institute. The Center for Molecular Medicine at Caltech is supported by the Gordon and Betty Moore Foundation. The operations at the SSRL are supported by the US Department of Energy (DOE) and the National Institutes of Health (NIH). GM/CA has been funded in whole or in part with federal funds from the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006). A portion of this research was supported by NIH grant U24GM129547 and performed at the PNCC at OHSU and accessed through EMSL (grid.436923.9), a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research. A.H. was supported by a Camille-Dreyfus Teacher Scholar Award (TC-15-082) and NIH grants R01-GM117360 and R01- GM111461, is an investigator of the Heritage Medical Research Institute (HMRI-15-09-01), and is a faculty scholar of the Howard Hughes Medical Institute (55108534). S.P. was supported by a predoctoral fellowship from the Boehringer Ingelheim Fonds and by an Amgen Graduate Fellowship through the Caltech-Amgen Research Collaboration. This research used resources of the Advanced Photon Source, a U.S. DOE Office of Science user facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
A version of this press release was originally published by Caltech.
The U.S. Department of Energy's APS at Argonne National Laboratory is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
