At the root of the devastating COVID-19 pandemic is the SARS-CoV-2 virus. One of the enzymes essential for this virus to function is PLpro, which is a protease that cleaves large viral proteins into smaller proteins. The proteolytic activity of this enzyme ultimately enables viral spread. In a recent study published by an international research team that carried out investigations at the U.S. Department of Energy’s Advanced Photon Source (APS), a chemical screen was done to identify irreversible inhibitors that are highly selective for PLpro. Using the APS high-brightness x-ray beams, the authors generated crystal structures of two particular potent inhibitors—VIR250 and VIR251—in complex with PLpro, allowing for a structural understanding of how these compounds mediate their inhibition. This work was published in the journal Science Advances.
The virus SARS-CoV-2 has a nanoscopic structure less than 200 nanometers in diameter. The SARS-CoV-2 virion consists of a lipid-enriched envelope encompassing a positive-sense single-stranded RNA genome. Viruses with a positive-sense RNA genome have the ability to utilize their genome as messenger RNA and directly translate viral proteins. The envelope protecting this genome is studded with proteins, the most prominent of which are spike proteins. The genome at the core of the SARS-CoV-2 virion contains close to 30,000 bases and encodes for various proteins that enable viral replication. More than 20 proteins originate from this genome, four of which are the spike, envelope, membrane, and nucleocapsid structural proteins.
One of these proteins is an enzyme called PLpro, which is a protease that breaks up larger proteins into smaller proteins. The smaller proteins freed from a larger polyprotein by PLpro ultimately enable viral replication by forming a replicase complex. Since this replicase complex is responsible for genome replication, PLpro is essential for viral replication. Given that SARS-CoV-2 is not infectious without this enzyme, PLpro has attracted considerable attention as a potential antiviral drug target.
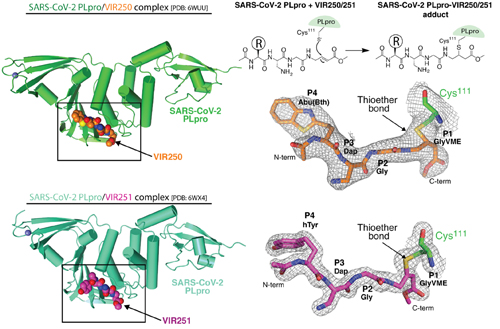
The authors began their study of PLpro by performing a comprehensive drug screen with the goal of identifying drugs that can irreversibly inhibit PLpro. Two of these chemicals—referred to as VIR250 and VIR251—displayed a high selectivity for different PLpro variants. Importantly, these compounds did not inhibit other similar human enzymes. Having identified these two potent compounds, the researchers determined the crystal structures of each compound in complex with PLpro (Fig. 1). This structural approach led the authors to identify how these inhibitors interact with and inhibit this enzyme. The authors additionally show that PLpro engages in deISGylating activity, which is thought to play an important role in evading a host's immune response. They demonstrate how this activity differs from SARS-CoV-2 and SARS-CoV-1, potentially contributing to our understanding of why the infectivity of these viruses is different. For the VIR250 and VIR251 crystals in complex with PLpro, structural data were collected at the Northeastern Collaborative Access Team beamline 24-ID-C at the APS.
With these potent inhibitors identified, the next step could be determining whether or not they can decrease mortality and attenuate deleterious symptoms in pre-clinical animal models. If so, they will represent viable therapeutics for increasing survival and improving the health of COVID-19 patients. — Alicia Surrao
See: Wioletta Rut1*, Zongyang Lv2, 3*, Mikolaj Zmudzinski1, Stephanie Patchett4, Digant Nayak2,3, Scott J. Snipas5, Farid El Oualid6, Tony T. Huang4**, Miklos Bekes7***‡, Marcin Drag1,5*****, Shaun K. Olsen2, 3****, “Activity profiling and crystal structures of inhibitor bound SARS-CoV-2 papain-like protease: A framework for anti–COVID-19 drug design,” Sci. Adv. 6: eabd4596 (16 October 2020). DOI: 10.1126/sciadv.abd4596
Author affiliations: 1Wroclaw University of Science and Technology, 2Medical University of South Carolina, 3University of Texas Health Science Center at San Antonio, 4New York University School of Medicine, 5Sanford Burnham Prebys Medical Discovery Institute, 6UbiQ Bio B.V., 7Independent Consultant
Correspondence: * [email protected], ** [email protected], , *** [email protected], **** [email protected], ***** [email protected] ‡Present address: Arvinas, Inc.
This project was supported by the National Science Center grant 2015/17/N/ST5/03072 (Preludium 9) in Poland (W.R.) and the “TEAM/2017-4/32” project, which is carried out within the TEAM program of the Foundation for Polish Science, co-financed by the European Union under the European Regional Development Fund (M.D.). W.R. is a beneficiary of a START scholarship from the Foundation for Polish Science. The Northeastern Collaborative Access Team beamlines are funded by the National Institute of General Medical Sciences from the National Institutes of Health (P30 GM124165). Research reported in this publication was supported by CPRIT RR200030 and NIH R01 GM115568 (S.K.O.), ES025166 (T.T.H.), and GM099040 (S.J.S.). Z.L. is a Hollings Cancer Center Postdoctoral Fellow, and S.P. is an American Cancer Society Postdoctoral Fellow (PF-18-235- 01-RMC). This research used resources of the APS, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract DE-AC02-06CH11357. Extraordinary facility operations were supported in part by the DOE Office of Science through the National Virtual Biotechnology Laboratory, a consortium of DOE national laboratories focused on the response to COVID-19, with funding provided by the Coronavirus CARES Act.
The U.S. Department of Energy's APS is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
