The original University of Bonn press release can be read here.
An international research team led by the University of Bonn has identified and further developed novel antibody fragments against the SARS coronavirus-2 (SARS-CoV-2). These "nanobodies" are much smaller than classic antibodies. The nanobodies therefore penetrate the tissue better and can be produced more easily in larger quantities. The researchers at the University Hospital Bonn have also combined the nanobodies into potentially particularly effective molecules. These attack different parts of the virus simultaneously. The approach could prevent the pathogen from evading the active agent through mutations. The results are published in the journal Science.
Antibodies are an important weapon in the immune system's defense against infections. They bind to the surface structures of bacteria or viruses and prevent their replication. One strategy in the fight against disease is therefore to produce effective antibodies in large quantities and inject them into patients. However, these antibodies have a complex structure, do not penetrate very deeply into the tissue, and may cause unwanted complications. Moreover, producing antibodies is difficult and time-consuming. They are, therefore, probably not suitable for widespread use.
"We focus on another group of molecules, the nanobodies," explains Florian Schmidt, who heads an Emmy Noether group on this promising new field of research at the University of Bonn's Institute of Innate Immunity. "Nanobodies are antibody fragments that are so simple that they can be produced by bacteria or yeast, which is less expensive."
However, the immune system produces an almost infinite number of different antibodies, and they all recognize different target structures. Only a very few of them are, for example, capable of defeating the SARS-CoV-2. Finding these antibodies is like searching for a single grain of sand on Germany's Baltic coast. "We first injected a surface protein of the coronavirus into an alpaca and a llama," Schmidt explains. "Their immune system then produces mainly antibodies directed against this virus. In addition to complex normal antibodies, llamas and alpacas also produce a simpler antibody variant that can serve as the basis for nanobodies."
A few weeks later, the researchers took a blood sample from the animals, from which they extracted the genetic information of produced antibodies. This "library" still contained millions of different construction plans. In a complex process, they extracted those that recognize an important structure on the surface of the coronavirus, the spike protein. "Altogether we obtained dozens of nanobodies, which we then analyzed further," explains Paul-Albert König, head of the Core Facility Nanobodies at the Medical Faculty of the University of Bonn and lead author of the study.
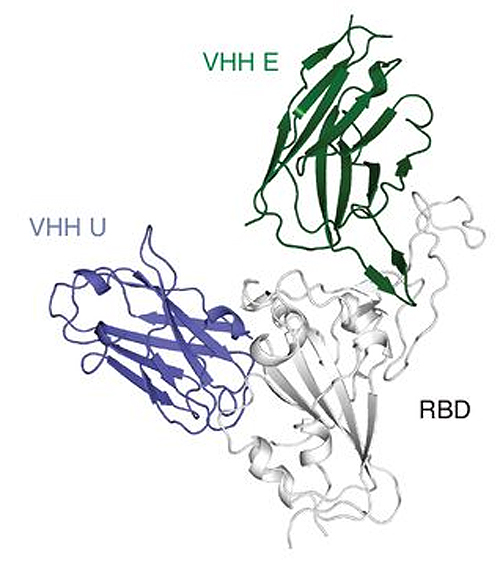
Four molecules actually proved to be effective against the pathogen in cell cultures. "Using x-ray structures and electron microscopy analyses, we were furthermore able to show how they interact with the spike protein of the virus," explains König. X-ray diffraction studies were carried out at the National Institute of General Medical Sciences and National Cancer Institute 23-ID-B x-ray beamline at the Advanced Photon Source, a DOE Office of Science user facility at Argonne National Laboratory; and beamline 12-1 of the Stanford Synchrotron Radiation Lightsource, a DOER Office of Science user facility at SLAC National Accelerator Laboratory. This work was done in the research groups around Martin Hällberg (Karolinska Institutet, Sweden) and Nicholas Wu (University of Illinois at Urbana-Champaign, USA) as well as Ian Wilson (Scripps Research Institute, USA). The spike protein is crucial for the infection: It acts like a velcro fastener with which the pathogen attaches to the attacked cell. Next, the velcro changes its structure: It discards the component that is important for attachment and mediates fusion of the virus envelope with the cell. "Nanobodies also appear to trigger this structural change before the virus encounters its target cell―an unexpected and novel mode of action," says König. "The change is likely to be irreversible; the virus is therefore no longer able to bind to host cells and infect them."
The researchers also exploit another major advantage of nanobodies over antibodies. Their simple structure allows straightforward combinations to form molecules that can be several hundred times more effective. "We have fused two nanobodies that target different parts of the spike protein," explains König. "This variant was highly effective in cell culture. Furthermore, we were able to show that this drastically reduces the probability of the virus becoming resistant to the active agent through escape mutations." The researchers are convinced that the molecules may be developed into a novel and promising therapeutic option.
Dioscure Therapeutics, a spin-off of the University of Bonn, will test the nanobodies in clinical studies. The success of the project is mainly based on the excellent cooperation of the participating research groups at the university with national and international cooperation partners, emphasizes Florian Schmidt, who is also a member of the Immunosensation2 Cluster of Excellence at the University of Bonn.
See: Paul-Albert König1,2*, Hrishikesh Das3, Hejun Liu4, Beate M. Kümmerer5,6, Florian N. Gohr2, Lea-Marie Jenster2, Lisa D.J. Schiffelers2, Yonas M. Tesfamariam2, Miki Uchima2, Jennifer D. Wuerth2, Karl Gatterdam7, Natalia Ruetalo8, Maria H. Christensen2, Caroline I. Fandrey2, Sabine Normann2, Jan M. P. Tödtmann1, Steffen Pritzl1, Leo Hanke9, Jannik Boos10, Meng Yuan4, Xueyong Zhu4, Jonathan Leo Schmid-Burgk11, Hiroki Kato12, Michael Schindler8, Ian A. Wilson4,13, Matthias Geyer7, Kerstin U. Ludwig10, B. Martin Hällberg3,14**, Nicholas C. Wu15,16,17***, and Florian I. Schmidt1,2****, “Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape,” Science 12 January 2021. DOI: 10.1126/science.abe6230
Author affiliations: 1Core Facility Nanobodies, Medical Faculty, University of Bonn, 2Institute of Innate Immunity, Medical Faculty, University of Bonn, 3Department of Cell and Molecular Biology, Karolinska Institutet, 4Department of Integrative Structural and Computational Biology, The Scripps Research Institute, 5Institute of Virology, Medical Faculty, University of Bonn, 6German Centre for Infection Research (DZIF), 7Institute of Structural Biology, Medical Faculty, University of Bonn, 8Institute for Medical Virology and Epidemiology, Section Molecular Virology, University Hospital Tübingen, 9Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, 10Institute of Human Genetics, Medical Faculty, University of Bonn, 11Institute for Clinical Chemistry and Clinical Pharmacology, Medical Faculty, University of Bonn, 12Institute of Cardiovascular Immunology, Medical Faculty, University of Bonn, 13The Skaggs Institute for Chemical Biology, The Scripps Research Institute, 14Centre for Structural Systems Biology (CSSB) and Karolinska Institutet VR-RÅC, 15Department of Biochemistry, University of Illinois at Urbana-Champaign, 16Carl R. Woese Institute for Genomic Biology, University of Illinois at Urbana-Champaign, 17Center for Biophysics and Quantitative Biology, University of Illinois at Urbana-Champaign
Correspondence: *[email protected], **[email protected], ***[email protected], ****[email protected]
In Germany, the study was financially supported by the Medical Faculty of the University of Bonn, the German Research Foundation, the Klaus Tschira Boost Funds, the Federal Ministry of Education and Research, the Baden-Württemberg Foundation and the MWK Baden-Württemberg. In the USA, the Bill and Melinda Gates Foundation, the U.S. Department of Energy, the National Institutes of Health (NIH), the National Institute of General Medical Sciences (NIGMS) and the National Cancer Institute (NCI) funded the project, in Sweden the Swedish Research Council and the Knut and Alice Wallenberg Foundation. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (P30GM133894). This research used resources of the Advanced Photon Source, an Office of Science User Facility operated for the U.S. DOE Office of Science by the Argonne National Laboratory under contract no. DE-AC02-06CH11357.
The U.S. Department of Energy's APS is one of the world’s most productive x-ray light source facilities. Each year, the APS provides high-brightness x-ray beams to a diverse community of more than 5,000 researchers in materials science, chemistry, condensed matter physics, the life and environmental sciences, and applied research. Researchers using the APS produce over 2,000 publications each year detailing impactful discoveries, and solve more vital biological protein structures than users of any other x-ray light source research facility. APS x-rays are ideally suited for explorations of materials and biological structures; elemental distribution; chemical, magnetic, electronic states; and a wide range of technologically important engineering systems from batteries to fuel injector sprays, all of which are the foundations of our nation’s economic, technological, and physical well-being.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC, for the U.S. DOE Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
