 Ebola virus is a member of the Ebolavirus genus, which also contains other pathogenic viruses such as Sudan virus and Bundibugyo virus. Ebola virus infection causes Ebola virus disease, a severe and frequently fatal hemorrhagic fever in humans and non-human primates that has implications for global biosecurity and public health. A major challenge in treating this disease lies in the fact that five different viruses in the Ebolavirus genus can cause it.
Ebola virus is a member of the Ebolavirus genus, which also contains other pathogenic viruses such as Sudan virus and Bundibugyo virus. Ebola virus infection causes Ebola virus disease, a severe and frequently fatal hemorrhagic fever in humans and non-human primates that has implications for global biosecurity and public health. A major challenge in treating this disease lies in the fact that five different viruses in the Ebolavirus genus can cause it.
This deadly illness cost the international community more than $50 billion to fight during the 2014–2016 West Africa outbreak alone. As of March 6, 2019, the ongoing EVD outbreak in the Democratic Republic of the Congo had infected nearly 1000 people, with an approximately 57% fatality rate.
Although these two most recent outbreaks were caused by Ebola virus itself, other viruses in the Ebolavirus genus have caused deadly outbreaks among humans, and likely will do so again in the future. However, because rapid identification of causative ebolavirus species can be difficult, treatment strategies and therapeutics are needed that can combat multiple virus species in the Ebolavirus genus. Such antibodies that can neutralize all five viruses are exceedingly rare.
In order to enter and infect the host animal, Ebola virus has to first attach to a host cell. A glycoprotein, termed GP, on the surface of the virus is critical for this attachment, and all five ebolaviruses have a single but different GP on each of their surfaces. Antibodies against these GPs are therefore attractive candidates for broadly reactive therapeutics that may be capable of treating EVD, regardless of the specific virus responsible for the disease outbreak.
Understanding the molecular interactions between broadly neutralizing antibodies and the Ebola virus GP is therefore crucial to the design of next-generation immunotherapeutic cocktails. Scientists from The Scripps Research Institute have recently focused on understanding how antibodies directed against ebolavirus GPs can mount an effective neutralization response against multiple members of the Ebolavirus genus. Using x-ray crystallography, the team collected diffraction data at the U.S. Department of Energy’s Advanced Photon Source (APS), in order to examine the three-dimensional molecular interaction between an antibody and its target, the ebolavirus GP. The results of this study will instruct vaccine design aimed at eliciting a broadly neutralizing humoral immune response against ebolaviruses.
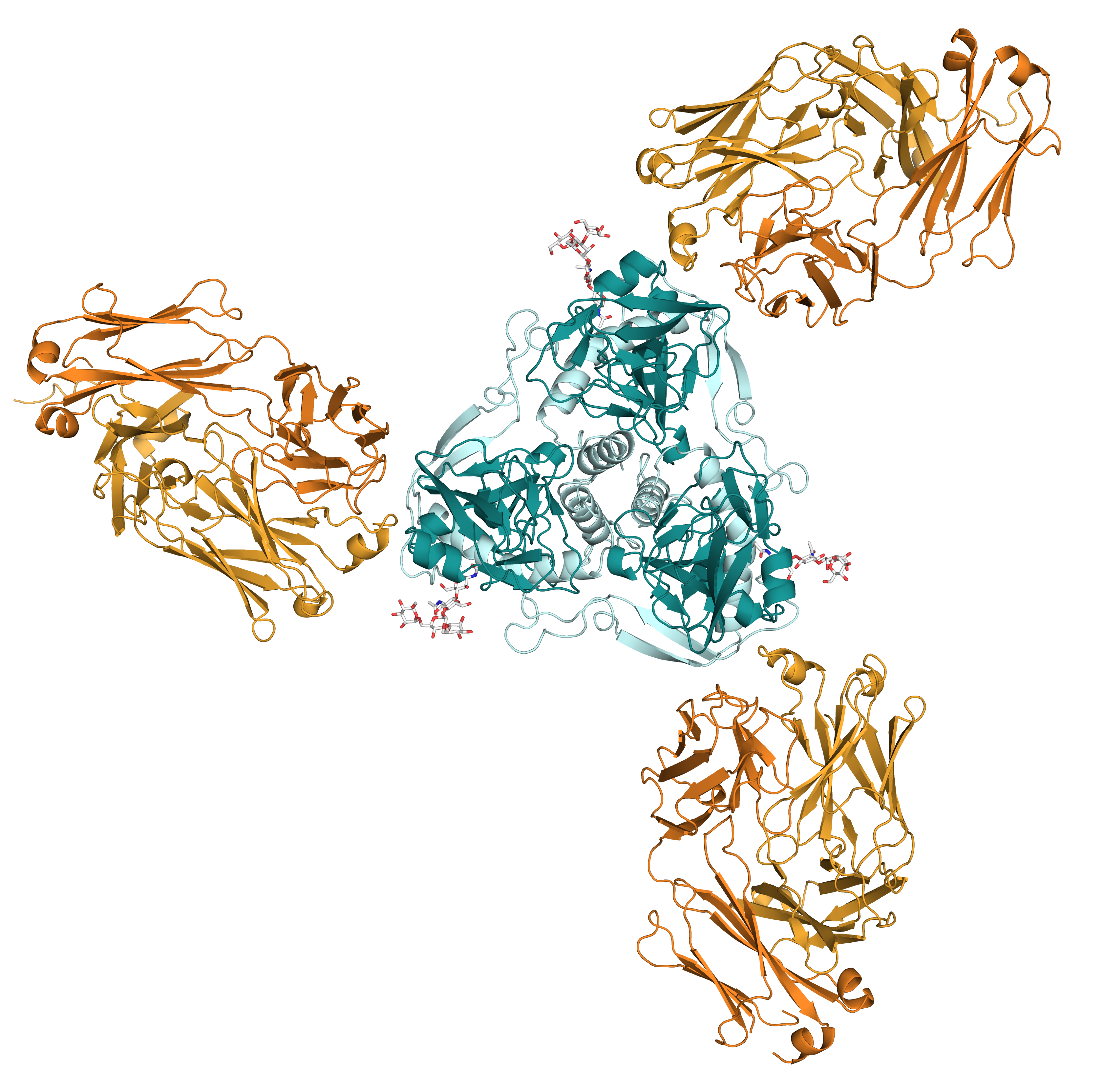
In this study, the researchers investigated how one broadly neutralizing antibody, ADI-15946, interacts with the viral glycoprotein. Working at the National Institute of General Medical Sciences and National Cancer Institute (GM/CA-XSD) structural biology facility x-ray beamline 23-ID-B at the APS, they evaluated a three-dimensional molecular model of the structure of both the viral GP epitope (the part that antibody binds to) and the antibody’s paratope (the part that binds to the epitope) in complex with the viral GP cellular receptor (Fig. 1). (The APS is an Office of Science user facility at Argonne National Laboratory.)
In particular, the model of the paratope enabled them to engineer the antibody to improve binding and neutralization of more distant members of the Ebolavirus genus. Functional insights provided by the study’s accompanying biochemistry also confirmed the structural observations of this interaction. Together, the data highlight an epitope on the various ebolavirus GPs that is susceptible to broad neutralization by a single antibody.
Unexpectedly, the researchers also found that the antibody’s binding site on the GP occupies a pocket on the GP surface that is usually closed off by a loop that descends from the glycan cap, or stalk region, of the GP.
Future ebolavirus vaccine design will need to take this observation into consideration in order to design an effective immunogen that is capable of eliciting broadly neutralizing antibodies that target the ADI-15946 paratope via a humoral immune response. ― Nicola Parry
See: Brandyn R. West, Crystal L. Moyer, Liam B. King, Marnie L. Fusco, Jacob C. Milligan, Sean Hui, and Erica Ollmann Saphire*, “Structural Basis of Pan-Ebolavirus Neutralization by a Human Antibody against a Conserved, Yet Cryptic Epitope,” mBio 9(5), e01674-18 (September/October 2018). DOI: 10.1128/mBio.01674-18.
Author affiliation: The Scripps Research Institute
Correspondence: * [email protected]
The authors acknowledge a National Institutes of Health (NIH) NIAID Centers of Excellence in Translational Research award for the Viral Hemorrhagic Fever Immunotherapeutic Consortium (U19 AI109762) and NIH R01 AI132204 for support. GM/CA-XSD is funded in whole or in part with Federal funds from the National Cancer Institute (ACB-12002) and the National Institute of General Medical Sciences (AGM-12006). This research used resources of the Advanced Photon Source, a U.S. Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC for the U.S. Department of Energy's Office of Science.
The U.S. Department of Energy's Office of Science is the single largest supporter of basic research in the physical sciences in the United States and is working to address some of the most pressing challenges of our time. For more information, visit the Office of Science website.
