DNA damage is a fact of life. On any given day, an organism’s DNA will suffer between 10,000 and 1,000,000 breaks or other damage. These problems are repaired by enzymes in our cells that fix the breaks, remove errors, and maintain the integrity of the genome. One of these DNA repair enzymes acts as a kind of molecular scissors to cut DNA at damage points and resolve tangles that can form when things go wrong. This must be done with great specificity in order to restore the DNA code to its previous state and not generate mutations.
Researchers from the Pohang University of Science and Technology (South Korea) and Argonne National Laboratory utilized macromolecular crystallography at the U.S. Department of Energy’s Advanced Photon Source (APS) at Argonne National Laboratory and the Pohang Accelerator Laboratory in South Korea to solve the structure of one of these molecular scissor proteins, called Mus81-Eme1, in a complex with DNA. Their analysis reveals the elegant way the enzyme recognizes, positions, and cuts the DNA, and then secures the two ends to avoid a new tangle. Their work furthers our understanding of the mechanisms of DNA repair, with positive implications for fields such as reproduction, cancer, and aging.
Most genes that have been found to affect lifespan are involved in DNA repair. If breaks and other DNA damage are not repaired, disease can result from adverse effects on normal gene expression and cell division functions that are critical to life.
One type of enzyme involved in DNA repair is a kind of molecular scissors called an endonuclease. Mus81-Eme1 is one of a family of structure-selective endonucleases responsible for resolution of complex DNA overlaps that are the result of DNA damage. These overlaps, or flaps, come in two different flavors, 5′ and 3′ (pronounced “5-prime” and “3-prime” in DNA parlance) depending on the direction of the DNA break that occurred.
The researchers in this study were interested in why the Mus81-Eme1 endonuclease specifically recognizes only the 3′ flaps and not the 5′ flaps.
In order to study this, they crystallized Mus81-Eme1 in complex with small pieces of DNA with either 3′ or 5′ flaps and collected diffraction data for the crystals at the Structural Biology Center Collaborative Access Team x-ray beamline 19-ID-D at the APS, and at the Pohang Accelerator Laboratory.
As one might expect for an enzyme that performs an important basic DNA repair function, Mus81-Eme1 works with elegance and precision. The team compared structures for Mus81-Eme1 in complex with one 5′ flap DNA and two 3′ flap DNAs. The 5′ flap structure showed that the DNA could bind to Mus81-Eme1 but the binding did not result in cutting of the DNA. Analysis of the 3′ flap structures revealed why.
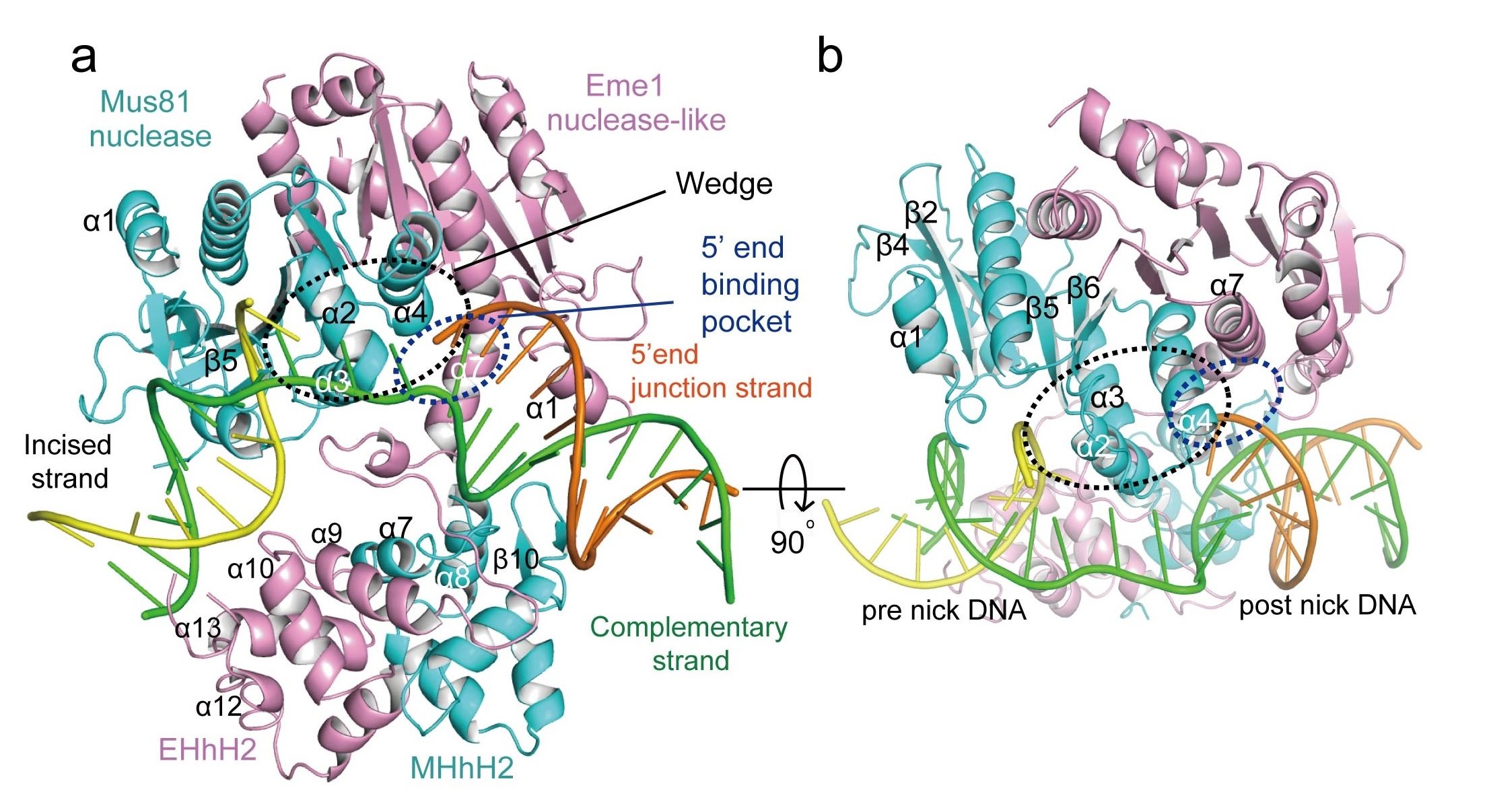
When Mus81-Eme1 binds to the 3′ flap DNAs it undergoes a dramatic shape change that converts it from a compact structure to a more open structure (see the figure). This unmasks a protein domain that acts as a wedge to keep the two cut ends separated. The end that is not being cut, the 5′ end, is held in place by a specialized binding pocket that keeps this end away from the action.
Next, Mus81-Eme1 bends the DNA to position the 3′ end right in the active site. Biochemically, the 5′ flaps don’t work because the free 3′ ends don’t fit properly into the binding pocket and so the DNA doesn’t bend to place the 5′ end into the active site.
The team validated their model in two ways. First, they confirmed reduced enzyme activity in mutant versions of Mus81-Eme1 based on structure model predictions. Second, in experiments in which they labeled both ends of the DNA with fluorescent tags, the addition of Mus81-Eme1 resulted in the two ends of the DNA moving closer together, supporting the bending formation that had been observed in the structure.
What’s next?
Because endonucleases like Mus81-Eme1 are responsible for repair of some of the most complex types of DNA damage, when these are not repaired due to a malfunction of an endonuclease, serious developmental and genetic disease can occur. The next step for the team will be to study the structures of other members of this endonuclease family, including one involved in a disease called Fanconi anemia, in order to provide a framework for curing these diseases.
— Sandy Field
See: Gwang Hyeon Gwon1, Aera Jo1, Kyuwon Baek1, Kyeong Sik Jin1, Yaoyao Fu1, Jong-Bong Lee1, YoungChang Kim2, and Yunje Cho1*, “Crystal structures of the structure-selective nuclease Mus81-Eme1 bound to flap DNA substrates,” EMBO J. 33 (9), 937 (2014). DOI:10.1002/embj.201487820
Author affiliations: 1Pohang University of Science and Technology, 2Argonne National Laboratory
Correspondence: * [email protected]
This work was supported by grants from the National R&D Program for Cancer Control, Ministry for Health and Welfare (1020280), National Research Foundation of Korea grant funded by the Korea government (MEST, No. 2012004028, No. 2012-054226, and No. 20120008833), a rising star program (POSTECH), and the BK21 program (Ministry of Education). The Structural Biology Center Collaborative Access Team is funded by the U.S. Department of Energy (DOE) Office of Biological and Environmental Research. This research used resources of the Advanced Photon Source, a U.S. Department of Energy Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under Contract No. DE-AC02-06CH11357.
Argonne National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.