The mechanism by which cells respond to stimuli and trigger hormonal responses, as well as the senses of sight, smell, and taste, has for the first time been brought into focus with the help of high-brightness x-rays provided by the U.S. Department of Energy Office of Science’s Advanced Photon Source (APS) at Argonne National Laboratory. This breakthrough will pave the way to new research avenues in drug discovery, cell signaling, and cellular regulation.
Receiving a signal, interpreting it, and responding correctly: these are three activities that cells must be good at in order to respond to stimuli. For responding to hormones, and for the senses of sight, smell, and taste, the activated receptors are coupled with G proteins. These G protein-coupled receptors (GPCRs) are involved in a complex set of steps in which an extracellular hormone or neurotransmitter binds to and activates a GPCR in the cell membrane. The activated receptor passes on the signal to a G-protein inside the cell, triggering reactions that ultimately create a cellular response to the stimulus.
The intricacy of this set of reactions has been appreciated—and studied—for some time. What was lacking, however, was a finely-detailed crystal structure of the GPCR as it actually signals across the membrane. The importance of acquiring this knowledge and knowing a lot more about how GPCRs function is apparent when we consider that the human genome contains over 800 GPCR genes.
The structure of protein receptors that are involved in cellular responses to stimuli is now available thanks to experiments carried out at the National Institute of General Medical Sciences and National Cancer Institute Collaborative Access Team (GM/CA-CAT) beamline 23-ID-B at the APS, and the process of cell signaling has come into crystal clear focus. This crystal structure is the first high-resolution look at transmembrane signaling by a GPCR and adds critical insight about signal transduction across the plasma membrane.
The general model for GPCR signaling is that the G-protein is activated by a receptor that has received a stimulus from outside the cell. The research team, comprising members from Stanford University, the University of Copenhagen, the University of Michigan, the University of Wisconsin, Vrije Universiteit Brussel, and Trinity College studied a specific model system for GPCR signaling that has long been used by biochemists and about which much was already known. In this system, a β2 adrenergic receptor (β2AR) is bound by the outside stimulus, known as an agonist, and then activates Gs, the stimulatory G protein for adenylyl cyclase.
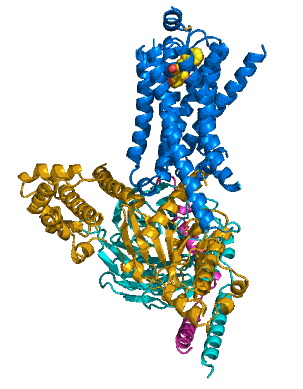
The researchers were able to capture the β2AR bound to the Gs protein before the latter went on to the next step in the dance, which would be binding a nucleotide. One technical challenge, and perhaps the reason that the crystal structure had been so elusive to date, was to create a stable β2AR-Gs complex in detergent solution. Once this problem was solved, the team could proceed to unveiling the crystal structure of this active-state β2AR-Gs complex (see the accompanying figure).
The new data allowed the research team to construct the early stages of GPCR signaling. Using their own observations, and other data collected on this model system, a mechanism for the initial stages of complex formation came into being. The data provided a pinpoint determination of exactly which changes in the molecules permitted that binding stage to carry on to completion.
Shape changes in the Gs, particularly in the nucleotide-binding pocket, prepared the protein for the next step. In the β2AR, the researchers were able to identify major changes in two areas of the molecule that fit well with understanding of how the β2AR interacts with molecules outside the cell and how the two molecules—β2AR and Gs—interact each other.
Article co-author Brian Kobilka, of Stanford University, recalled how the project reached this important milestone: “For the past five years, Roger Sunahara and I have been working together to understand how receptors and G proteins interact with each other. This was the next logical step after getting the first inactive-state structures of the β2AR alone in 2007.
“The receptor and G protein purification procedures were well established at the time we started the project, but most of the other methods were developed during the project. The minibeam and the rastering‡ capabilities of the GM/CA-CAT beamline at the APS were essential for collecting the diffraction data on these crystals.”
Future research, Kobilka said, involves “how the complex forms and dissociates after activation.” — Mona Mort
See: Søren G. F. Rasmussen1,2, Brian T. DeVree3, Yaozhong Zou1, Andrew C. Kruse1, Ka Young Chung1, Tong Sun Kobilka1, Foon Sun Thian1, Pil Seok Chae4, Els Pardon5, Diane Calinski3, Jesper M.Mathiesen1, Syed T. A. Shah6, Joseph A. Lyons7, Martin Caffrey6, Samuel H. Gellman4, Jan Steyaert5,Georgios Skiniotis7,William I. Weis1, Roger K.Sunahara3**, and Brian K. Kobilka1*, “Crystal structure of the b2 adrenergic receptor–Gs protein complex,” Nature, advance online publication, 19 July 2011. DOI: 10.1038/nature10361
Author affiliations: 1Stanford University School of Medicine, 2University of Copenhagen, 3University of Michigan Medical School, 4University of Wisconsin, Madison, 5Vlaams Instituut voor Biotechnologie, 6Trinity College, 7University of Michigan
Correspondence: * [email protected] or ** [email protected]
The authors acknowledge support from National Institutes of Health Grants NS028471 (B.K.K.) and GM083118 (B.K.K. and R.K.S.), GM56169 (W.I.W.), P01 GM75913 (S.H.G), and P60DK-20572 (R.K.S.), GM75915, P50GM073210 and U54GM094599 (M.C.), Science Foundation Ireland (07/IN.1/B1836) and FP7 COST Action CM0902 (M.C.),the Mathers Foundation (B.K.K. and W.I.W.), the Lundbeck Foundation (Junior Group Leader Fellowship, S.G.F.R.), the University of Michigan Biomedical Sciences Scholars Program (R.K.S.), the Fund for Scientific Research of Flanders (FWO-Vlaanderen) and the Institute for the encouragement of Scientific Research and Innovation of Brussels (ISRIB) (E.P. and J.S.), and The Danish Council for Independent Research, Medical Sciences (J.M.M.).
GM/CA-CAT has been funded in whole or in part with Federal funds from the National Cancer Institute (Y1-CO-1020) and the National Institute of General Medical Sciences (Y1-GM-1104).
Use of the Advanced Photon Source at Argonne National Laboratory was supported by the DOE’s Office of Science under Contract No. DE-AC02-06CH11357.
See also: “Cell signalling: It's all about the structure,” by Lizzie Buchen, naturenews, 24 Augusts, 2011. DOI:10.1038/476387a
‡Rastering: A technique used in protein crystallography, wherein the sample being studied is stepped across an x-ray beam center in a systematic grid pattern, and an x-ray exposure is taken at each of the grid locations. A demonstration video can be seen and heard here: https://www.gmca.aps.anl.gov/
The Advanced Photon Source at Argonne National Laboratory is one of five national synchrotron radiation light sources supported by the U.S. Department of Energy’s Office of Science to carry out applied and basic research to understand, predict, and ultimately control matter and energy at the electronic, atomic, and molecular levels, provide the foundations for new energy technologies, and support DOE missions in energy, environment, and national security. To learn more about the Office of Science x-ray user facilities, visit http://science.energy.gov/user-facilities/basic-energy-sciences/.
Argonne National Laboratory seeks solutions to pressing national problems in science and technology. The nation's first national laboratory, Argonne conducts leading-edge basic and applied scientific research in virtually every scientific discipline. Argonne researchers work closely with researchers from hundreds of companies, universities, and federal, state and municipal agencies to help them solve their specific problems, advance America's scientific leadership and prepare the nation for a better future. With employees from more than 60 nations, Argonne is managed by UChicago Argonne, LLC for the U.S. Department of Energy's Office of Science.