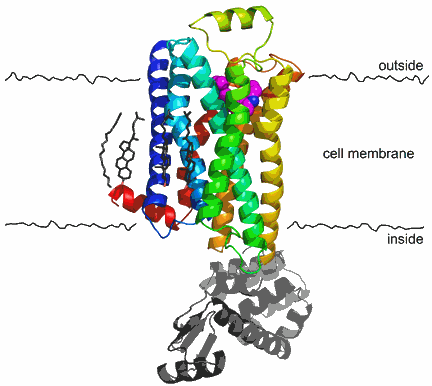
More than 40 years after beta blockers were first used clinically, scientists can finally got a close-up look at the drugs' molecular target: the β2-adrenergic receptor. The work is particularly exciting because it offers the first glimpse into an important, but scientifically elusive family of human proteins called G protein-coupled receptors (GPCRs).
More than half of all drugs given to patients work by targeting one or another GPCR, found on body cells, to steer the cell's machinery toward healing an illness. Researchers from Stanford University and the Scripps Research Institute have determined what the β2-adrenergic receptor looks like at the molecular level, giving them the keys to greater control of the process.
With more than 300 members, GPCRs constitute the largest family of proteins found in the membranes of cells. These cellular receptors function like molecular switches to promote or stifle a multitude of biological processes within the cells. GPCRs play a critical role in heart disease, blood pressure regulation, inflammation and psychological disorders. The research promises not only to speed the discovery of new and improved drugs, but also to broaden our understanding of human health and disease.
Published online October 21 as an Advance Online Publication in Nature and in the October 25 issue of Science Express, the research was supported by the National Institutes of Health.
The work represents a technical tour de force that required the scientists to devise several new techniques. Many of the difficulties arose because the receptor is a membrane protein-one of the trickiest molecules to capture in three-dimensional detail.
After considerable efforts with the protein in a natural form, the researchers, led by Brian Kobilka of Stanford University, turned to protein engineering. To overcome problems with the protein's floppiness, they replaced part of the protein with another, stiffer molecule, essentially clamping the protein into place so they could work with it more easily. They also utilized several new methods to minimize the amount of the protein needed for detailed structural studies.
After years of effort to produce high-quality crystals, the researchers also needed an especially narrow and brilliant X-ray beam to examine crystals that were very small and very sensitive to radiation damage. They used a 6-micron mini-beam developed this year at the General Medicine and Cancer Institutes Collaborative Access Team (GM/CA-CAT) at the Advanced Photon Source. Data collection took place at GM/CA CAT beamline 23ID-B, and earlier at the European Synchrotron Radiation Facility (ESRF), beamlines ID-13 and ID23-2.
"This is an absolutely remarkable advance," said Jeremy M. Berg, Ph.D., director of NIGMS which, in addition to supporting the innovative work of both academic laboratories, also plays a leading role in support of GM/CA CAT. "Many laboratories around the world are trying to reveal the secrets of these proteins and this new structure takes this field to a new level."
See: Søren G. F. Rasmussen, Hee-Jung Choi, Daniel M. Rosenbaum, Tong Sun Kobilka, Foon Sun Thian, Patricia C. Edwards, Manfred Burghammer, Venkata R. P. Ratnala, Ruslan Sanishvili, Robert F. Fischetti, Gebhard F. X. Schertler, William I. Weis & Brian K. Kobilka, “ Crystal structure of the human beta2 adrenergic G-protein-coupled receptor,” doi:10.1038/nature06325. Nature 15 November 2007 450: 383-387. Epub 21 October 2007.
See: Daniel M. Rosenbaum, Vadim Cherezov, Michael A. Hanson, Søren G. F. Rasmussen, Foon Sun Thian, Tong Sun Kobilka, Hee-Jung Choi, Xiao-Jie Yao, William I. Weis, Raymond C. Stevens, and Brian K. Kobilka, “GPCR Engineering Yields High-Resolution Structural Insights into β2-Adrenergic Receptor Function,” doi: 10.1126/science.1150609. Science 23 November 2007 318: 1266-1273. Epub 25 October 2007.
Contact: Brian Kobilka, [email protected]
The original Stanford University press release can be found here: http://news-service.stanford.edu/news/2007/october31/med-protien-103107.html
The original NIH press release can be found here: https://www.nih.gov/news/pr/oct2007/nigms-29.htm
This work was supported by NIH grant F32 GM082028 (to D.M.R.); the Lundbeck Foundation (to S.G.F.R); NIH Roadmap Initiative grant P50 GM073197 and Protein Structure Initiative P50 GM62411 (to R.C.S.); NINDS grant NS028471, the Mather Charitable Foundations, Lundbeck, and the NIH Roadmap Initiative grant R21 GM075811 (to B.K.K.). H.-J.C. and W.I.W. were supported in part by NIH grant R01 GM056169. GM/CA CAT is funded by the National Cancer Institute (Y1-CO-1020) and the National Institute of General Medical Sciences (Y1-GM-1104). Use of the Advanced Photon Source was supported by the U. S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357.
Argonne is a U.S. Department of Energy laboratory managed by UChicago Argonne, LLC